Function Over Form:
Phenotypic Integration and the Evolution of the Mammalian Skull
By Claire Asher, on 8 December 2014
Our bodies are more than just a collection of independent parts – they are complex, integrated systems that rely upon precise coordination in order to function properly. In order for a leg to function as a leg, the bones, muscles, ligaments, nerves and blood vessels must all work together as an integrated whole. This concept, known as phenotypic integration, is a pervasive characteristic of living organisms, and recent research in GEE suggests that it may have a profound influence on the direction and magnitude of evolutionary change.
Phenotypic integration explains how multiple traits, encoded by hundreds of different genes, can evolve and develop together such that the functional unit (a leg, an eye, the circulatory system) fulfils its desired role. Phenotypic integration could be complete – every trait is interrelated and could show correlated evolution. However, theoretical and empirical data suggest that it is more commonly modular, with strong phenotypic integration within functional modules. This modularity represents a compromise between a total lack of trait coordination (which would allow evolution to breakdown functional phenotypic units) and the evolutionary inflexibility of complete integration. Understanding phenotypic integration and its consequences is therefore important if we are to understand how complex phenotypes respond to natural selection.
It is thought that phenotypic integration is likely to constrain evolution and render certain phenotypes impossible if their evolution would require even temporary disintegration of a functional module. However, integration may also facilitate evolution by coordinating the responses of traits within a functional unit. Recent research by GEE academic Dr Anjali Goswami and colleagues sought to understand the evolutionary implications of phenotypic integration in mammals.
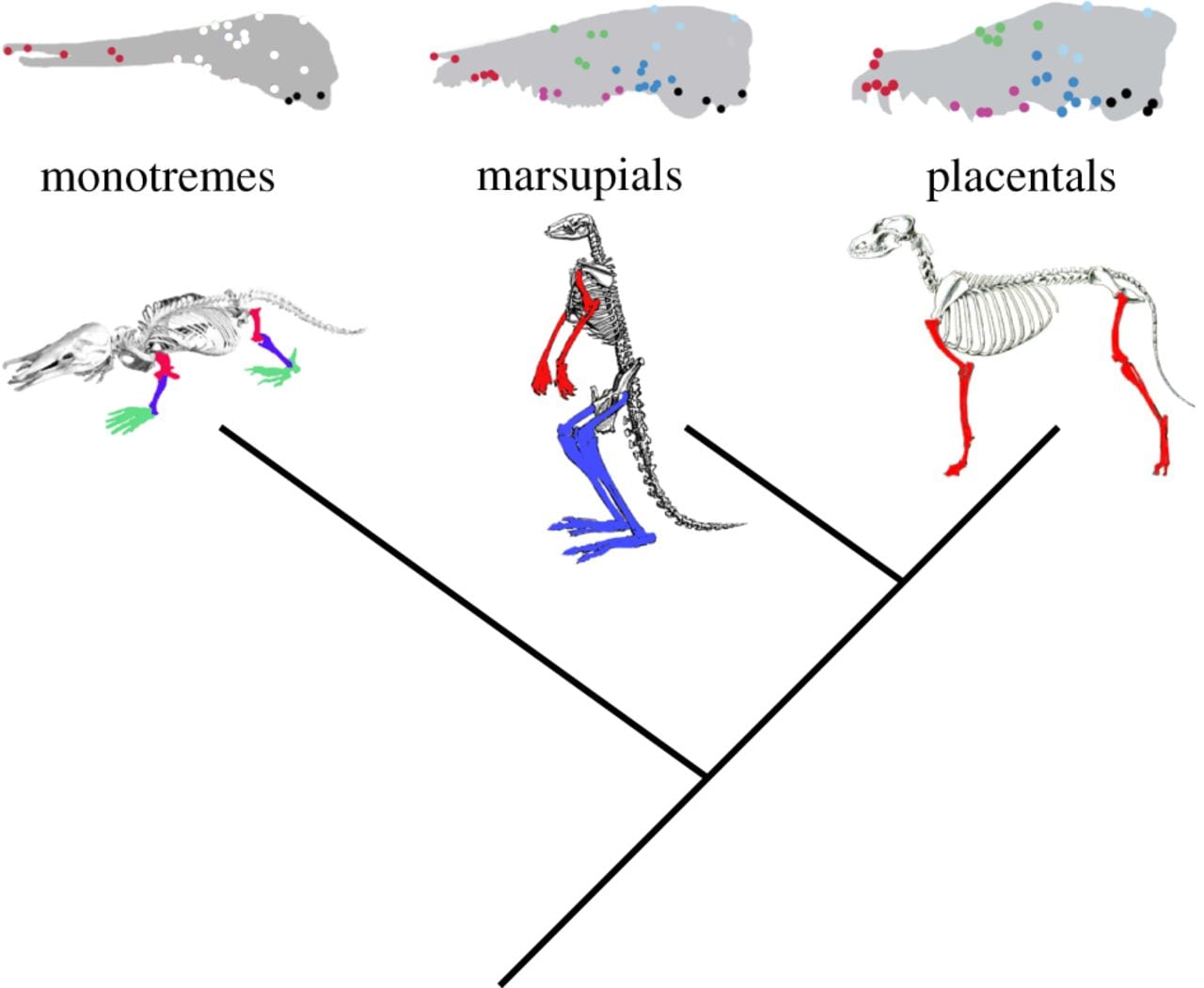
Expanding on existing mathematical models, and applying these to data from 1635 skulls from nearly 100 different mammal species including placental mammals, marsupials and monotremes, Dr Goswami investigated the effect of phenotypic integration on evolvability and respondability to natural selection. Comparing between a model with two functional modules in the mammalian skull and a model with six, the authors found greater support for a larger number of functional modules. Monotremes, whose skulls may be subject to different selection pressures due to their unusual life history, did not fit this pattern and may have undergone changes in cranial modularity during the early evolution of mammals. Compared with random simulations, real mammal skulls tend to be either more or less disparate from each other, suggesting that phenotypic integration may both constrain and facilitate evolution under different circumstances. The authors report a strong influence of phenotypic integration on both the magnitude and trajectory of evolutionary responses to selection, although they found no evidence that it influences the speed of evolution.
 Thus, phenotypic integration between functional modules appears to have a profound impact on the direction and extent of evolutionary change, and may tend to favour convergent evolution of modules that perform the same function (e.g bird and bat wings for powered flight), by forcing individuals down certain evolutionary trajectories. The influence of phenotypic integration on the speed, direction and magnitude of evolution has important implications for the study of evolution, particularly when analysing fossil remains, since it can make estimates of the timing of evolutionary events more difficult. Failing to incorporate functional modules into models of evolution will likely reduce their accuracy and could produce erroneous results.
Thus, phenotypic integration between functional modules appears to have a profound impact on the direction and extent of evolutionary change, and may tend to favour convergent evolution of modules that perform the same function (e.g bird and bat wings for powered flight), by forcing individuals down certain evolutionary trajectories. The influence of phenotypic integration on the speed, direction and magnitude of evolution has important implications for the study of evolution, particularly when analysing fossil remains, since it can make estimates of the timing of evolutionary events more difficult. Failing to incorporate functional modules into models of evolution will likely reduce their accuracy and could produce erroneous results.
Phenotypic integration is what holds together functional units within an organism as a whole, in the face of natural selection. Modularity enables traits to evolve independently when their functions are not strongly interdependent, and prevents evolution from disintegrating functional units. Through these actions, phenotypic integration can constrain or direct evolution in ways that might not be predicted based on analyses of traits individually. This can have important impacts upon the speed, magnitude and direction of evolution, and may tend to favour convergence.
Original Article:
This research was made possible by support from the Natural Environment Research Council (NERC), and the National Science Foundation (NSF).
 Close
Close